I have previously studied how specialised chromatin domains, namely telomeres, are spatially organised within the nucleus by sumoylation. Current projects involve studying the role of chromatin remodelling enzymes in genome stability and their interplay with telomeres.
Previous research projects
SUMO regulates telomere localisation in S. cerevisiae
SUMO (Small Ubiquitin-like modifier) is a post-translational modification added onto lysine residues and has been linked to genome stability processes. We observed that deletion of the PIAS-like SUMO E3 ligase SIZ2, but not other E3 ligases, led to telomere delocalisation from the nuclear periphery. We showed that Siz2 promotes both telomere anchoring pathways of budding yeast (Sir4 and Yku70/80) and sumoylates these proteins in vivo. Importantly, we showed that the defect in Yku70 and Yku80-4 mediated chromatin anchoring in siz2Δ cells can be overcome by making a linear fusion of Yku70 or Yku80 to SUMO. Deletion of SIZ2 also resulted in a telomerase-dependent increase in telomere length. Remarkably, the localisation and length phenotypes seen in siz2Δ cells appear to be functionally linked. We found that critically short telomeres detach from the nuclear periphery when they are most efficiently elongated. Our study was the first to show an effect of sumoylation on telomere anchoring and argues that Siz2-mediated control of telomere position helps regulate stable telomere maintenance. 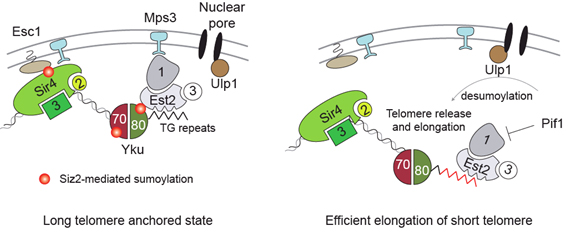
Figure 1. Model of Siz2 mediated effects on telomere localisation and elongation (Ferreira et al NCB 2011)
Characterising C. elegans telomere localisation
We have extended our work in yeast by looking at telomere localisation within the multicellular animal, C. elegans. Using a telomere FISH protocol that we optimised (Figure 2), we find that C. elegans telomeres are preferentially bound to the nuclear periphery. This perinuclear association increases during embryogenesis and persists in later larval stages. Remarkably, we find that, as in yeast, PIAS-mediated sumoylation is required for telomere anchoring in C. elegans, being dependent on the SUMO E3 ligase GEI-17. We show that telomere position is independent of the Ku complex and of H3 K9 methylation but instead is dependent on the shelterin component POT-1 anchoring at nuclear envelope to SUN-1. To our knowledge, this is the first characterisation of telomere localisation and anchoring in C. elegans.
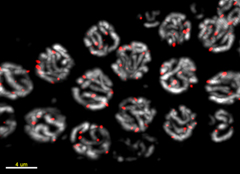
Figure 2. Telomere (red) and DNA (grey) staining of pachytene cells within the C. elegans germline (Ferreira et al JCB 2013).
Ongoing research projects
Chromatin remodelling and genome stability
ATP-dependent chromatin remodeling enzymes (Snf2-family ATPases) are a large class of motor proteins that have increasingly been shown to be important for DNA repair. Additionally, mutations in a number of Snf2 proteins lead to various developmental disorders as well as carcinogenesis in humans. However, despite their importance, the mechanism by which many Snf2 proteins promote genome stability remains poorly understood. This represents not only a lack of understanding of a fundamental biological process but also a lost opportunity to manipulate genome stability processes for medical benefit.
There is significant interplay between Snf2-family proteins and the sumoylation pathway. Many chromatin remodelling enzymes are SUMO modified, and chromatin remodelling enzymes are required to mediate SUMO-directed transcriptional repression. Understanding the role of sumoylation in Snf2 function may provide an important means to modulate their function.
Using C. elegans to understand ATRX
Mutation of ATRX in humans causes alpha-thalassemia, mental retardation (ATRX) syndrome and is also linked to a sub-type of cancers characterised by the alternative lengthening of telomeres (ALT) pathway. We are studying XNP-1, the worm homolog of ATRX, to better understand the connections between the developmental and telomeric roles of ATRX.
Protection of telomeric overhangs
Telomeres are nucleoprotein complexes that protect the ends of chromosomes, thereby preventing DNA damage and genome instability. Interestingly, chromosome ends are found as single-stranded DNA (ssDNA). The G-rich strand of telomeric ssDNA can form secondary structures, including guanine quadruplexes (G4s), that influence telomere stability. To regulate this, all organisms encode Protection of Telomere (POT) proteins. We have been characterising the POT proteins in C. elegans to understand how they work together to maintain genome stability.